
Imputation of low-pass genotypes using pedigree-based methods
Compte rendu Stage M2
1 Apr, 2026
Méthodes de génotypage
- Restricted number of loci
- Low error rate
- Low cost
- low coverage (<3X)
- Missing data
- Low cost
- Exhaustive
- Expensive
- Low number of samples
Low-pass \(\rightarrow\) Requires imputation algorithms
Du séquencage aux Variant Calling File (.vcf)
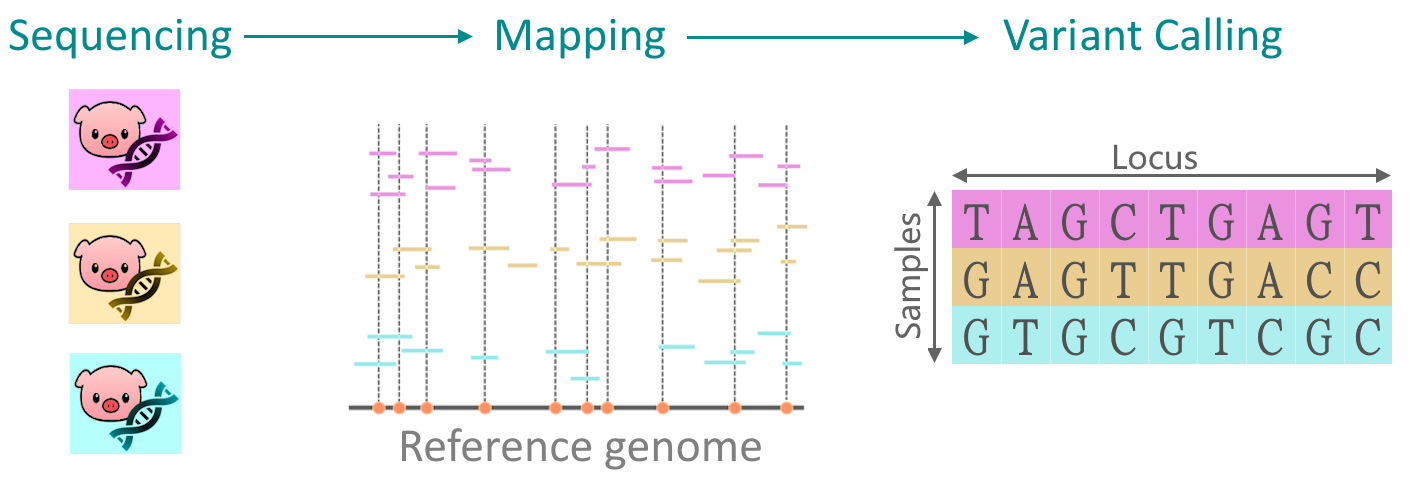
Vraisemblance de chaque génotypes
Penetrance: Distribution de la vraisemblance de chaque génotypes
| \(u_i\) | \(0\) | \(1\) | \(2\) |
|---|---|---|---|
| \(P(D|G)\) | 0.01 | 0.23 | 0.81 |
Dans les fichiers de variant calling (.vcf)
Dans un .vcf on utilise le “Phredscore Likelihood” (PL)
\[ PL = -10 * \log{P(D | G)} \]
Imputation
- Low-pass: Penetrance bruitée \(\rightarrow\) Imputation:
- 2 méthodes existantes : LD-based et Pedigree-based (mendel)
Méthodes basées sur le LD
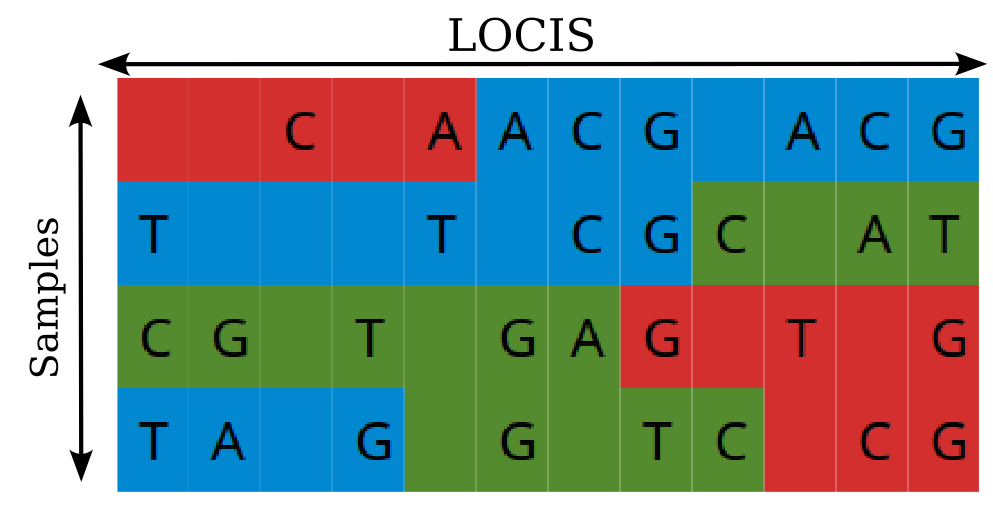
- Stitch (Davies et al. 2017)
- Glimpse (Rubinacci et al. 2021)
Imputation
Méthodes basées sur le pedigree
Pedigree
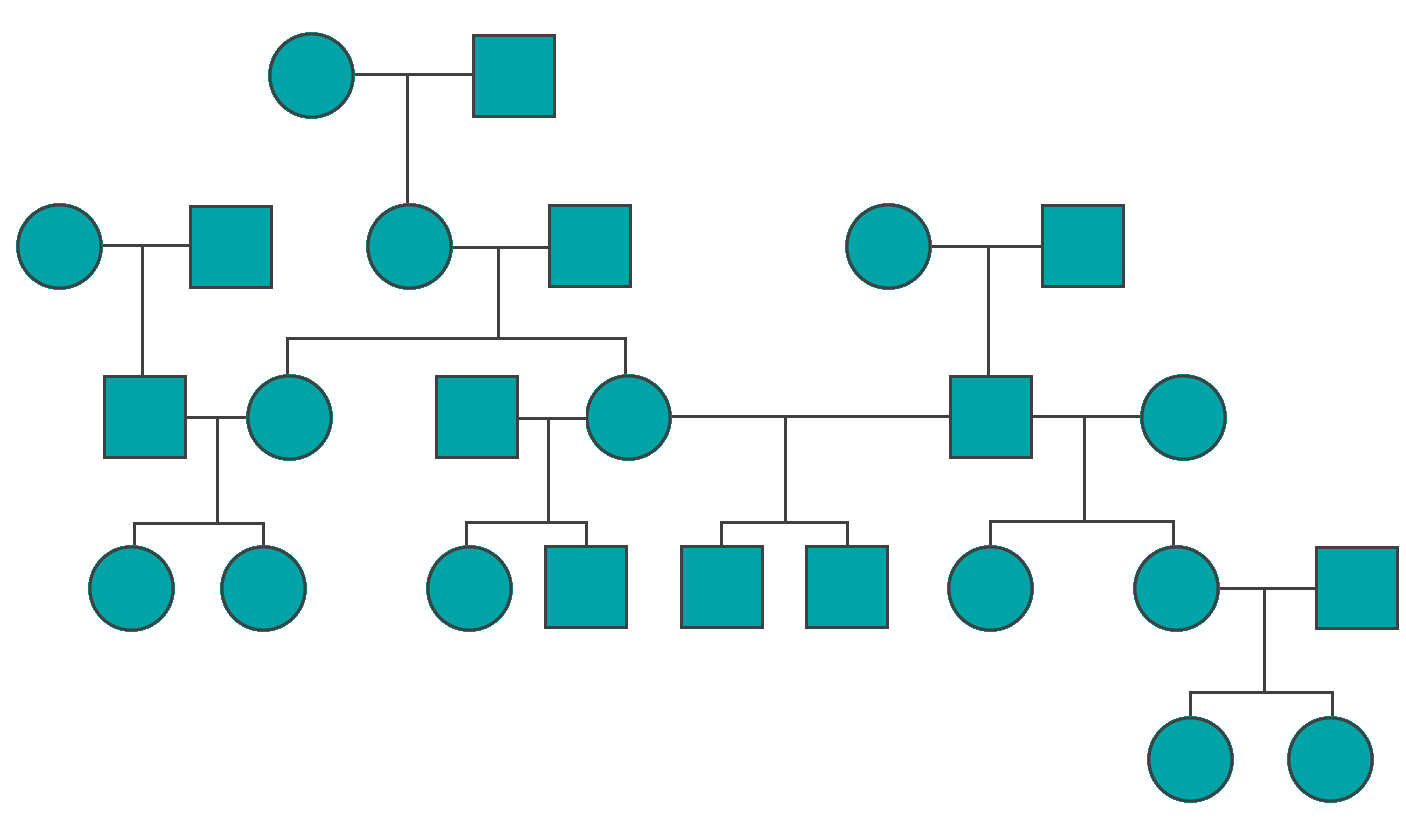
Exemples de méthodes
- FImpute
- Alphapeel
Imputation
Estimation de \(P(u_i|Y) \rightarrow\) Peeling
Peeling
Formule Fernando et al.
\[ \textcolor{orange}{Pr(u_i|y)} \propto \textcolor{red}{a_i(u_i)}\textcolor{green}{g(y_i|u_i)}\textcolor{blue}{\prod p_{ij}(u_i)} \]
Avec:
- \(\textcolor{green}{g(y_i|u_i)}\) ~ \(P(D|G)\)
- \(\textcolor{red}{a_i(u_i)} \rightarrow\) Anterior
- \(\textcolor{blue}{ p_{ij}(u_i)} \rightarrow\) Posterior
Peeling
Formule Fernando et al.
fonction Posterior
\[ \begin{align} \textcolor{blue}{p_{ij}(u_i)} = \sum_{u_j}\Bigg[\textcolor{red}{a_j(u_j)}\textcolor{green}{g(y_j|u_j)}\textcolor{blue}{\prod_{\stackrel{k\in S_j}{k\ne i}}p_{jk}(u_j)} \\ \times \prod_{k\in C_{ij}}\Bigg[\sum_{u_k}\textcolor{magenta}{tr(u_k|u_i ,u_j)}\textcolor{green}{g(y_k|u_k)} \textcolor{blue}{\prod_{l \in S_k}p_{kl}(u_k)}\Bigg]\Bigg] \end{align} \]
Peeling
Formule Fernando et al.
fonction Anterior
\[ \begin{align} \textcolor{red}{a_i(u_i)} =\sum_{u_m}\Bigg\{ \textcolor{red}{a_m(u_m)}\textcolor{green}{g(y_m|u_m)}\textcolor{blue}{\prod_{\stackrel{j\in S_m}{j\ne f}}p_{mj}(u_m)} \\ \times \sum_{u_f}\Big\{\textcolor{red}{a_f(u_f)}\textcolor{green}{g(y_f|u_f)}\textcolor{blue}{\prod_{\stackrel{j\in S_f}{j\ne m}}p_{fj}(u_f)}\\ \times \textcolor{magenta}{tr(u_i|u_m,u_f)} \\ \times \prod_{\stackrel{j\in C_mf}{j\ne i}}\Big[\sum_{u_j}\textcolor{magenta}{tr(u_j|u_m ,u_f)}\textcolor{green}{g(y_i|u_i)} \textcolor{blue}{\prod_{k \in S_i}p_{kj}(u_k)}\Big]\Big\}\Bigg\} \end{align} \]
Peeling
Aspect récursif
- Récursivité : fonctions qui s’appellent elles-mêmes
- Conditions d’arrêts:
- posterior: Individus sans enfants \(\rightarrow\) 1
- anterior: Fondateurs \(\rightarrow\) \(\hat{f}\)
Inconvénient majeur
Boucles récursivité impossible
Iterative Peeling
- L’algorithme iteratif (Kerr et Kinghorn, 1996.)
- pro: pedigree avec boucles
- con: \(\widehat{P(u_i|Y)}\)
Fonctionnement
- Initialisation des antérieurs et posterieurs
- Repeat :
- Peel down
- mise à jours des anteriors
- Peel up
- mise à jours des posteriors
- Peel down
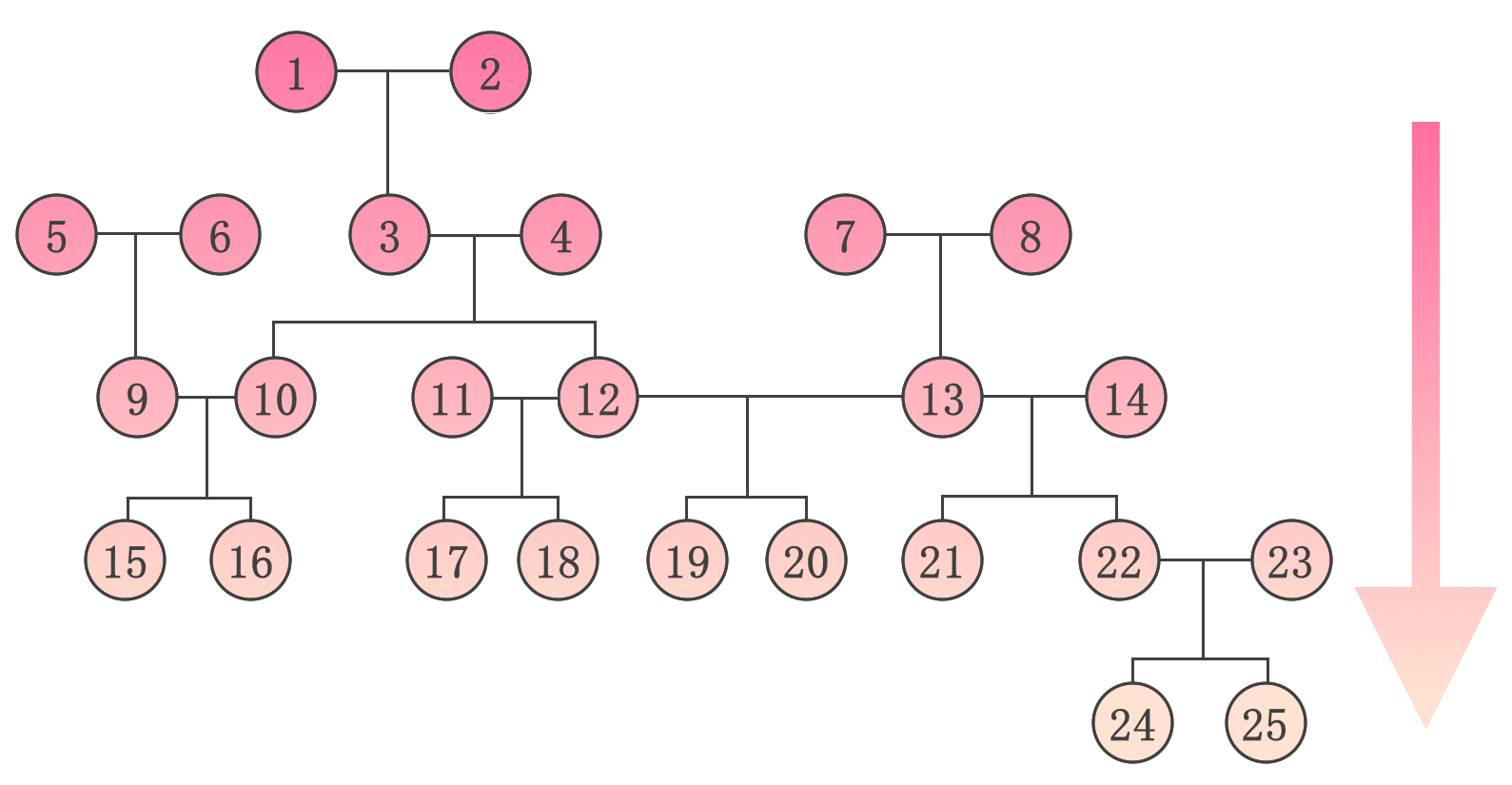
Iterative Peeling
- L’algorithme iteratif (Kerr et Kinghorn, 1996.)
- pro: pedigree avec boucles
- con: \(\widehat{P(u_i|Y)}\)
Fonctionnement
- Initialisation des antérieurs et posterieurs
- Repeat :
- Peel down
- mise à jours des anteriors
- Peel up
- mise à jours des posteriors
- Peel down
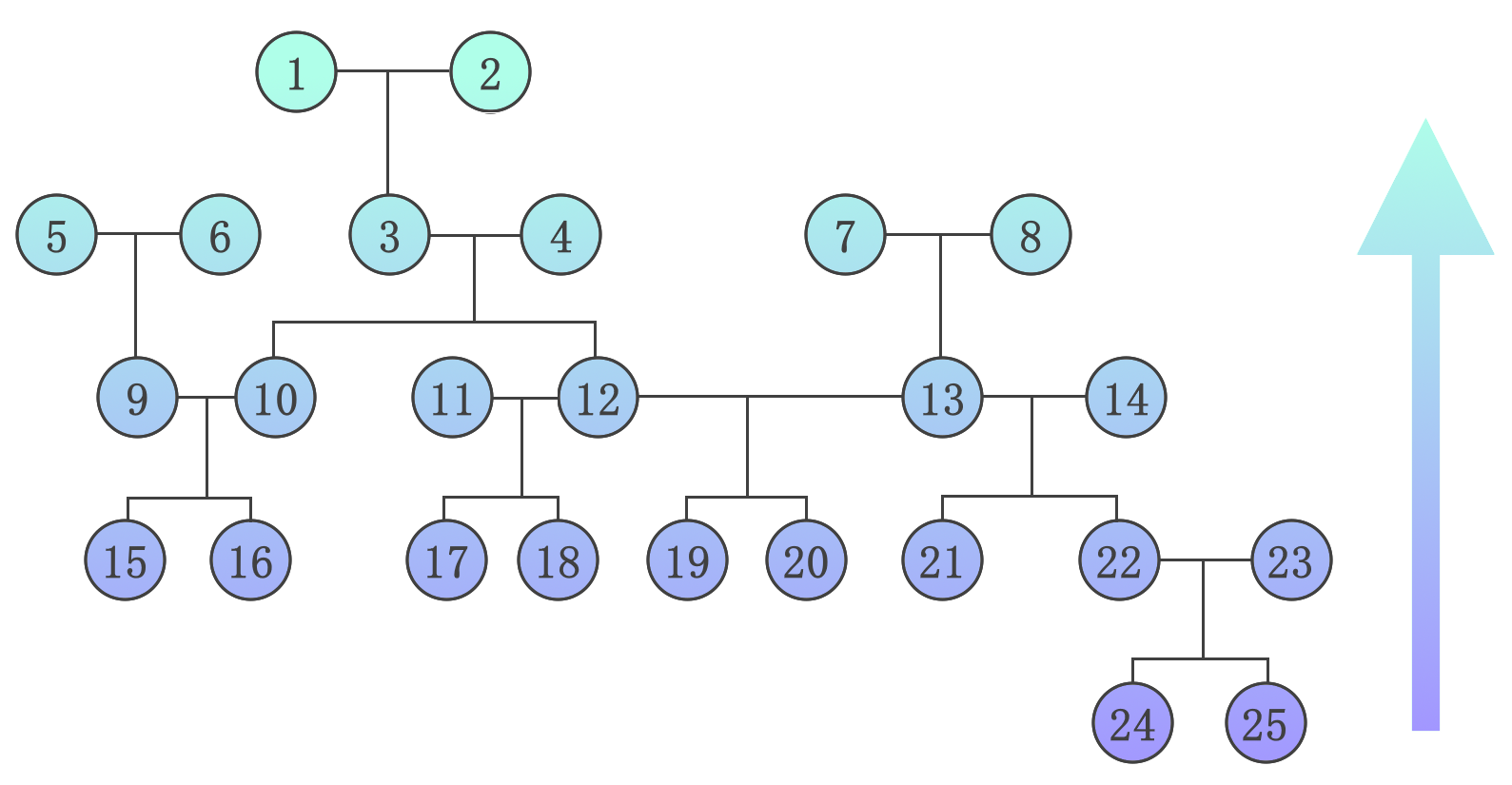
Implémentation
- Outil yapp
- Python
- calcul scientifique scipy
- gestion de fichiers .vcf panda
- stockage de données zarr
- gestion de projet Git
- travail en log
- scaling
- démonstration
- implémentation
Données simulées pedigree + genotypes: SimuPOP
Jeux de données
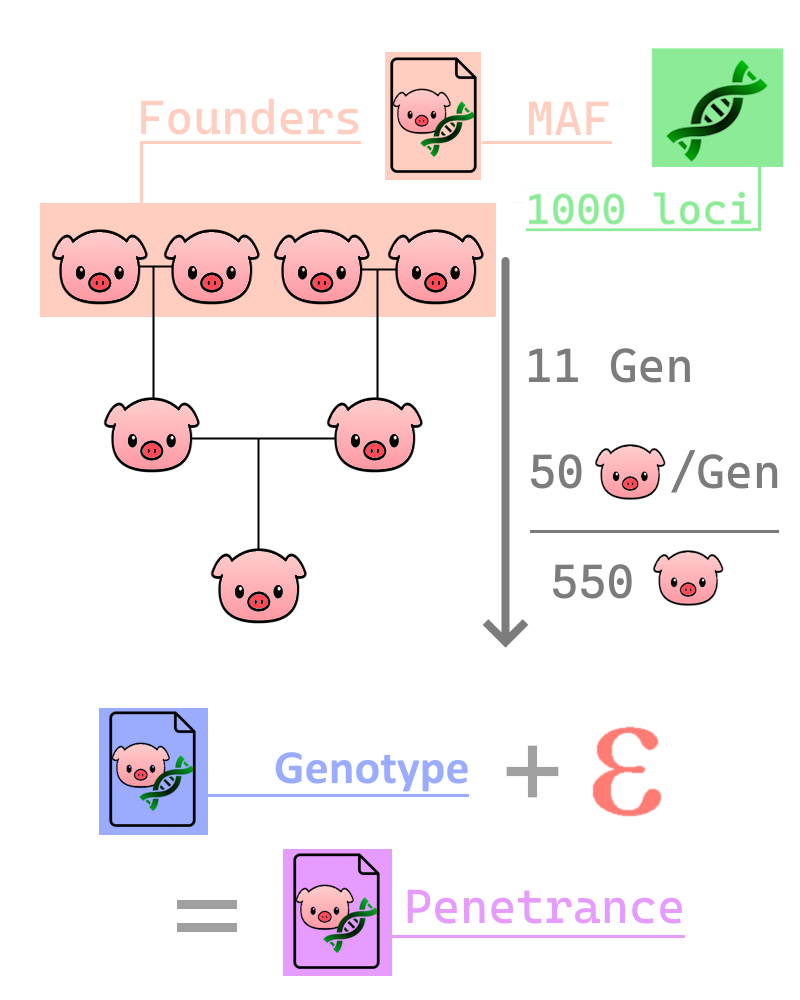
Simulation
- Génotype des fondateurs générés (algo coalescence)
- Générer un Pedigree (random mating)
- Génotype des individus générés
- Transmission Mendélienne
- loci indépendants
- Simulation de Penetrance Table 2
| Observed \(y_i\) | Formula |
|---|---|
| 0 (AA) | \((1-\epsilon)^2\) |
| 1 (Aa) | \(2 \cdot (1-\epsilon) \cdot \epsilon\) |
| 2 (aa) | \(\epsilon^2\) |
| NA | \(1\) |
Problème numérique Scaling
Problème: \(a_i(u_i)\) et \(p_{ij}(u_i)\) \(\simeq 0\)
Solution:
Scaling: \[ \begin{cases} a_i(u_i)=C_i \centerdot a_i^* (u_i) \\ g_i(y_i|u_i)=E_i \centerdot g_i^*(y_i|u_i) \\ p_{ij}(u_i) = D_{ij} \centerdot p_{ij}^*(u_i) \end{cases} \]
- On a donc: \[ \textcolor{Tan}{Pr(u_i|y)}\propto a_i^* (u_i) g_i^*(y_i|u_i) \prod_j p_{ij}^*(u_i) \]
Test d’estimation de MAF
Taux d’erreur \(\epsilon = 10^{-4}\)
Taux d’erreur \(\epsilon = 10^{-1}\)

Qualité d’estimation (\(\epsilon = 10^{-1}\))

Distribution de densité (\(\epsilon = 10^{-1}\))

Simulation de données low-pass
Pour un tableau de génotypes connus \(G\) \((I,L)\)
Simuler \(P(Y = \{\textcolor{Emerald}{D},\textcolor{Apricot}{R}\}|G)\):
Pour chaque \(i,l\) avec \(l \in L\) loci et \(i \in I\) individus:
- Tirer une profondeur \(\textcolor{Emerald}{D_{i,l}}\) où \(\textcolor{Emerald}{D_{i,l}} \sim \text{Poisson}(D_{\text{moy}})\)
- Tirer \(\textcolor{Apricot}{R_{i,l}}\) reads où \(\textcolor{Apricot}{R_{i,l}} \sim \text{Binomial}(\textcolor{Emerald}{D_{i,l}},P = f(G_{i,l},\textcolor{BrickRed}{\epsilon}))\)
- \(P(Y_{i,l} = \{\textcolor{Emerald}{D_{i,l}},\textcolor{Apricot}{R_{i,l}}\}|G_{i,l})= \begin{cases} 0: & \textcolor{BrickRed}{\epsilon}^{\textcolor{Apricot}{R_{i,l}}} \quad (1-\textcolor{BrickRed}{\epsilon})^{\textcolor{Emerald}{D_{i,l}} - \textcolor{Apricot}{R_{i,l}}} \\ 1: & \frac{1}{2}^\textcolor{Emerald}{D_{i,l}} \\ 2: & (1- \textcolor{BrickRed}{\epsilon})^{\textcolor{Apricot}{R_{i,l}}} \quad \textcolor{BrickRed}{\epsilon}^{\textcolor{Emerald}{D_{i,l}} - \textcolor{Apricot}{R_{i,l}}} \end{cases}\)
- scaling
| \(G_{i,l}\) | \(0/0\) | \(0/1\) | \(1/1\) |
|---|---|---|---|
| \(f(G_{i,l},\textcolor{BrickRed}{\epsilon})\) | \(\textcolor{BrickRed}{\epsilon}\) | \(1\over2\) | \(1 - \textcolor{BrickRed}{\epsilon}\) |
Données réelles (PorcQTL)
Donnée low-pass + array
- Évaluation sur Genotype array (Ground truth)
- \(P(u_i|Y)\) estimé sur du low-pass \(l \in L_{\text{array}} \cap L_{\text{low-pass}}\)
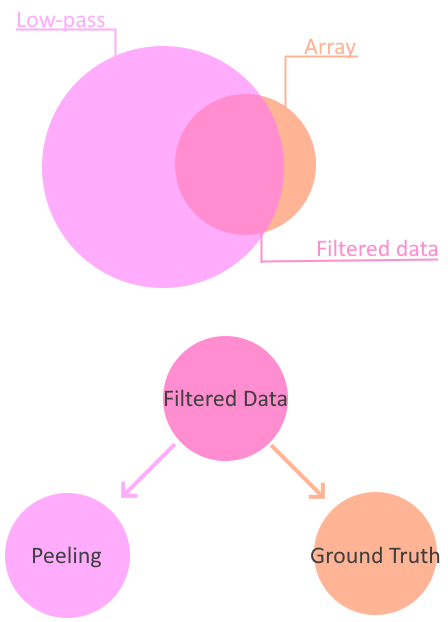
Applications
Peeling avec des données low-pass:
- Imputation :
- du génotype avec \(P(u_i|Y)\)
- locus manquants
- individus non génotypés
- Single-locus: \(\rightarrow\) besoin de méthodes utilisant LD
Assignation de parenté
Tester des parents candidats par rapport à un parent imputé
- Imputer le génotype du parent manquant
- Identifier le parent le plus probable

Perspectives du projet
- Test Genomewide
- Implémentation dans yapp
- Parallélisation
- Benchmark
- Alphapeel
- Glimpse
- Stitch
Slide Bonus!
Le code potentiel de génération de Penetrance
from scipy import stats
import numpy as np
random_state = 43
N, L, Dm, epsilon = 10, 1000, 5, 0.01
G = np.random.choice((0,1,2), size = (L,N), replace=True)
D = stats.poisson.rvs(Dm,size = (L,N), random_state = random_state)
pf = np.array((epsilon,0.5,1-epsilon))
pf_G = pf[G.ravel()].reshape((L,N))
R1 = stats.binom.rvs(D,pf_G, random_state = random_state)
penetrance = np.stack([
R1 * np.log(epsilon) + (D-R1) * np.log(1-epsilon),
D*np.full((L,N),np.log(0.5)),
R1*np.log(1-epsilon) + (D-R1) * np.log(epsilon)
],axis=-1)
penetrance = np.einsum("lnu->lun",penetrance)
Pied de page